Errata: More Than Life Itself
I fantasized the number of errors in More Than Life Itself to be zero, but hoped that it would be small, and that they would be slight and trivially fixable. My hopes were, fortunately, actualized. Tim Gwinn had found many of the errors (for which I am grateful), and he first posted his list on his website panmere.com. I also thank the readers who have been sending me corrections.
1 Praeludium
p.28 1.9 and write → and writes
p.36 l.1 The union should be intersection.
2 Principium
p.49 l.-8 a1,…,an → a1,…,an+1
p.49 l.-7 xn Ran y → xn Ran+1 y
p.56 2.30 Let f and g be observable of X. → Let f and g be observables of X.
p.57 2.30 (27): The final [x]Rg should be [x]Rf .
p.59 2.36 is and ordered pair → is an ordered pair
3 Continuatio
p.63 3.5 if it satisfy → if it satisfies
p.72 l.-8 Hesse diagrams → Hasse diagrams
p.73 3.27 Equivalently, lattice → Equivalently, a lattice
p.73 3.28 Equivalently, lattice → Equivalently, a lattice
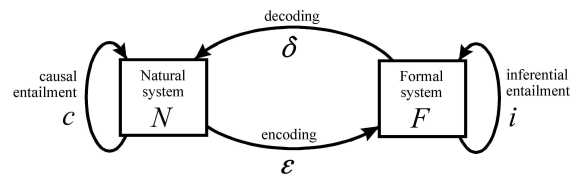
4 The Modelling Relation
p.97 l.-8 The encoding arrow α → The encoding arrow ε
5 Causation
p.107 l.12 restrictive too encompass → restrictive to encompass
pp.122–123 The second half of Section 5.16 has been rewritten.
(An overly stringent application of “a mapping uniquely determines its domain and co-domain” led to the disallowance of two modes of connection of two mappings. These restrictions are my oversight, and turn out to be unnecessary. I have since lifted them in a 2010 paper “Relational biology of symbiosis”, Axiomathes 20(4), 495-509.
springerlink.com/content/h481521x7264626l.
p.124 l.-2 because diagram (35) my → because diagram (35) may
p.130 l.14 If in they heart → If in thy heart
6 Topology
pp.140–141 Section 6.10 has been rewritten.
This is to allow the acceptance of the disallowed connections (see note on pp.122–123 above).
p.146 The xi in (30) is larger than the other symbols. Reduced.
p.141 6.11 and (b) G has an Eulerian circuit → and (b) G has an Eulerian path
p.147 ll.6–7 Changed tab position so that ‘compositions’ fits in line 6.
p.156 The beginning of Section 6.24 has been rewritten
to mention pseudographs.
p.157 ll.8–9 it would force (65) → it would force (66)
p.157 l.10 but (65) contradicts → but (66) contradicts
p.157 l.-4 requires the some, → requires that some,
p.158 l.2 (61) → (67)
p.158 l.5 .. → .
pp.159–160 Discussion following Theorem 6.28 has been rewritten,
and a new Section 6.29 has been added.
Solid-headed arrow self-loops are discussed in detail.
7 The Category of Formal Systems
p.170 l.3 s ∈ S → s ∈ S1
p.173 ll.6–7 Three occurrences: in S → in S1
p.174 l.-3 that if is a statement → that if Σ is a statement
p.182 l.7 does not → do not
p.184 l.16 headed arrows → headed arrow
p.184 l.21 relation diagram → relational diagram
8 Simple Systems
p.213 l.4 a corresponding subsequence of program lengths
→ a corresponding subsequence with program lengths
11 Living Systems
p.259 Bertalanffy quote Hierarchical order is found → [Hierarchical order] is found
The actual line is “A similar hierarchy is found…”, where the similarity refers to the subject of the previous sentence: “One attractive scheme of hierarchical order (there are others) is that of Boulding (Table 1.2).”
p.260 11.2 “Nothing comes from nothing.” → “Ex nihilo nihil fit.”
p.281 11.16 Traversability as a Relation Diagram
→ Traversability as a Relational Diagram
13 Ontogenic
p.313 l.-4 my → may
p.314 l.-5 deploring → deploying
Appendix Category Theory
p.344 l.-1 picture (1) → picture (2)
p.345 l.1 A-morphism commutativity → A-morphism associativity
p.355 l.2 an B-object → a B-object
Bibliography
p.376 l.12 [2000] ‘Autobiographical…’ → [2006] ‘Autobiographical…’
Index
p.381 col.2 ll.-6 332 → 332–333
p.385 col.1 ll.2–3 213–216–219 → 213–216, 219